# For display purposes:
from IPython.display import HTML
from itables import to_html_datatable
# For the simulation modelling and analysis:
import itertools
import os
import time
from joblib import Parallel, delayed, cpu_count
import matplotlib.colors as mcolors
import numpy as np
import pandas as pd
import plotly.express as px
import plotly.graph_objects as go
import scipy.stats as st
import simpy
from sim_tools.distributions import ExponentialLearning objectives:
- Recognise importance of sharing code for creation of tables and figures.
- View some example tables and figures.
Relevant reproducibility guidelines:
- STARS Reproducibility Recommendations (⭐): Include code to generate the tables, figures, and other reported results.
- STARS Reproducibility Recommendations: Save outputs to a file.
Pre-reading:
This page uses the results saved on the Scenario and sensitivity analysis page.
Required packages:
The following imports are required. These should be available from environment setup on the Structuring as a package page.
On this page, we have three new imports: os, matplotlib.colors and plotly.graph_objects.
library(dplyr)
library(ggplot2)
library(kableExtra)
library(knitr)When reporting results from a simulation, you will usually include tables and figures. To make your work reproducible, you should:
1. Make your tables and figures using code.
2. Share that code alongside your results.
This is just as important as sharing model and scenario code. In Heather et al. (2025), only one study included the code needed to generate all its reported results. Without this code for the others, recreating outputs was time-consuming and challenging. It required guesswork about which results to extract, how to clean them, and what processing steps were taken when producing the published tables and figures.
This page will show some examples of creating tables and figures, but there are many different ways you could describe and visualise your simulation results. The key takeaway from the page is simple: write code and share it!
Simulation results
This page will use the results saved on the Scenario and sensitivity analysis page.
Scenario analysis
scenario = pd.read_csv("scenarios_resources/python_scenario_results.csv")
HTML(to_html_datatable(scenario.head(200)))| Loading ITables v2.5.2 from the internet... (need help?) |
Sensitivity analysis
sensitivity = pd.read_csv("scenarios_resources/python_sensitivity_results.csv")
HTML(to_html_datatable(sensitivity.head(200)))| Loading ITables v2.5.2 from the internet... (need help?) |
Scenario analysis
scenario <- read.csv(file.path("scenarios_resources", "r_scenario_results.csv"))
kable(scenario) |> scroll_box(height = "400px")| X | replication | arrivals | mean_wait_time_doctor | mean_time_with_doctor | utilisation_doctor | mean_queue_length_doctor | mean_time_in_system | mean_patients_in_system | scenario | interarrival_time | number_of_doctors |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 1 | 8 | 0.0000000 | 10.626767 | 0.4811191 | 0.0000000 | 5.909446 | 1.5730708 | 1 | 4 | 3 |
| 2 | 2 | 14 | 1.3489715 | 16.474475 | 0.8415543 | 1.4037854 | 6.529895 | 4.0585764 | 1 | 4 | 3 |
| 3 | 3 | 8 | 3.3185568 | 12.576986 | 0.9214641 | 0.6425483 | 14.293369 | 2.4214309 | 1 | 4 | 3 |
| 4 | 4 | 9 | 7.0789803 | 8.229096 | 1.0000000 | 1.9960925 | 14.810041 | 3.2462020 | 1 | 4 | 3 |
| 5 | 5 | 11 | 11.8431217 | 8.866916 | 1.0000000 | 2.9270728 | 16.007073 | 4.2730406 | 1 | 4 | 3 |
| 6 | 1 | 9 | 1.1561321 | 12.600360 | 0.7545527 | 0.3739683 | 6.972457 | 2.4937655 | 2 | 5 | 3 |
| 7 | 2 | 10 | 0.2185845 | 15.462225 | 0.8137894 | 0.7190226 | 7.019465 | 3.0715488 | 2 | 5 | 3 |
| 8 | 3 | 8 | 1.6612341 | 11.704709 | 0.8144131 | 0.2671798 | 10.033307 | 1.7999155 | 2 | 5 | 3 |
| 9 | 4 | 4 | 1.3862671 | 8.268402 | 0.5633356 | 0.9223766 | 8.714268 | 0.8490077 | 2 | 5 | 3 |
| 10 | 5 | 7 | 1.1383003 | 8.211288 | 0.8027837 | 0.1753468 | 8.159880 | 1.5179933 | 2 | 5 | 3 |
| 11 | 1 | 4 | 0.0000000 | 16.756011 | 0.5829234 | 0.0000000 | 7.594901 | 1.6885375 | 3 | 6 | 3 |
| 12 | 2 | 7 | 0.0000000 | 13.951263 | 0.6247704 | 0.0000000 | 9.878982 | 1.8347756 | 3 | 6 | 3 |
| 13 | 3 | 11 | 0.0000000 | 10.642901 | 0.6696266 | 0.0000000 | 5.471170 | 2.1742571 | 3 | 6 | 3 |
| 14 | 4 | 5 | 2.1255795 | 7.962361 | 0.4490071 | 0.2915664 | 10.087940 | 1.3321312 | 3 | 6 | 3 |
| 15 | 5 | 6 | 0.1698502 | 7.931855 | 0.6664404 | 0.0000000 | 7.985399 | 0.9107067 | 3 | 6 | 3 |
| 16 | 1 | 5 | 0.0000000 | 14.928869 | 0.5309547 | 0.0000000 | 8.986369 | 1.5562112 | 4 | 7 | 3 |
| 17 | 2 | 5 | 0.0000000 | 16.557442 | 0.4795871 | 0.0000000 | 9.466947 | 1.2655765 | 4 | 7 | 3 |
| 18 | 3 | 8 | 0.0000000 | 10.066787 | 0.5494173 | 0.0000000 | 3.468666 | 2.0459750 | 4 | 7 | 3 |
| 19 | 4 | 5 | 0.9021887 | 7.962361 | 0.4504371 | 0.3962059 | 8.864549 | 1.3634454 | 4 | 7 | 3 |
| 20 | 5 | 4 | 0.0000000 | 8.753199 | 0.3340787 | 0.0000000 | 7.926292 | 0.8053229 | 4 | 7 | 3 |
| 21 | 1 | 3 | 0.0000000 | 20.737360 | 0.4033890 | 0.0000000 | 8.986369 | 1.2630193 | 5 | 8 | 3 |
| 22 | 2 | 5 | 0.0000000 | 16.557442 | 0.4525148 | 0.0000000 | 9.466947 | 1.1847657 | 5 | 8 | 3 |
| 23 | 3 | 7 | 0.0000000 | 11.304930 | 0.4485586 | 0.0000000 | 3.882443 | 1.8976861 | 5 | 8 | 3 |
| 24 | 4 | 6 | 0.0000000 | 6.932281 | 0.3689745 | 0.0000000 | 4.751341 | 1.0837230 | 5 | 8 | 3 |
| 25 | 5 | 3 | 0.0000000 | 9.563759 | 0.2902283 | 0.0000000 | 7.926292 | 0.5974274 | 5 | 8 | 3 |
| 26 | 1 | 11 | 0.2873017 | 8.979263 | 0.7388989 | 0.0000000 | 5.072421 | 2.0511195 | 6 | 4 | 4 |
| 27 | 2 | 7 | 0.0000000 | 11.322431 | 0.5589778 | 0.0000000 | 5.722932 | 1.9471475 | 6 | 4 | 4 |
| 28 | 3 | 10 | 0.3279443 | 13.891435 | 0.8192434 | 0.0000000 | 11.073710 | 2.3708711 | 6 | 4 | 4 |
| 29 | 4 | 6 | 0.0000000 | 7.155858 | 0.3352238 | 0.0000000 | 6.212797 | 0.9218882 | 6 | 4 | 4 |
| 30 | 5 | 8 | 0.0000000 | 9.299128 | 0.3465470 | 0.0000000 | 7.062117 | 1.6075429 | 6 | 4 | 4 |
| 31 | 1 | 9 | 0.0000000 | 14.361497 | 0.5564580 | 0.0000000 | 5.909446 | 2.0694053 | 7 | 5 | 4 |
| 32 | 2 | 8 | 0.0000000 | 10.902873 | 0.5209432 | 0.0000000 | 5.040122 | 1.6635569 | 7 | 5 | 4 |
| 33 | 3 | 8 | 0.1002196 | 12.491221 | 0.7258143 | 0.0000000 | 8.233837 | 1.9542346 | 7 | 5 | 4 |
| 34 | 4 | 5 | 0.0000000 | 7.220270 | 0.3280956 | 0.0000000 | 8.605687 | 0.8995180 | 7 | 5 | 4 |
| 35 | 5 | 5 | 0.0000000 | 7.764737 | 0.2639238 | 0.0000000 | 7.764737 | 0.9782088 | 7 | 5 | 4 |
| 36 | 1 | 4 | 0.0000000 | 16.756011 | 0.4371926 | 0.0000000 | 7.594901 | 1.6885375 | 8 | 6 | 4 |
| 37 | 2 | 7 | 0.0000000 | 13.951263 | 0.4685778 | 0.0000000 | 9.878982 | 1.8347756 | 8 | 6 | 4 |
| 38 | 3 | 11 | 0.0000000 | 10.921073 | 0.5067605 | 0.0000000 | 5.471170 | 2.1742571 | 8 | 6 | 4 |
| 39 | 4 | 6 | 0.0000000 | 6.932281 | 0.3478921 | 0.0000000 | 6.932281 | 1.0897496 | 8 | 6 | 4 |
| 40 | 5 | 4 | 0.0000000 | 8.753199 | 0.2551105 | 0.0000000 | 7.926292 | 0.9878955 | 8 | 6 | 4 |
| 41 | 1 | 5 | 0.0000000 | 14.928869 | 0.3982160 | 0.0000000 | 8.986369 | 1.5562112 | 9 | 7 | 4 |
| 42 | 2 | 5 | 0.0000000 | 16.557442 | 0.3596903 | 0.0000000 | 9.466947 | 1.2655765 | 9 | 7 | 4 |
| 43 | 3 | 8 | 0.0000000 | 10.066787 | 0.4120630 | 0.0000000 | 3.468666 | 2.0459750 | 9 | 7 | 4 |
| 44 | 4 | 6 | 0.0000000 | 6.932281 | 0.3489646 | 0.0000000 | 6.932281 | 1.2656769 | 9 | 7 | 4 |
| 45 | 5 | 4 | 0.0000000 | 7.539044 | 0.2003486 | 0.0000000 | 5.772494 | 0.5339783 | 9 | 7 | 4 |
| 46 | 1 | 3 | 0.0000000 | 20.737360 | 0.3025417 | 0.0000000 | 8.986369 | 1.2630193 | 10 | 8 | 4 |
| 47 | 2 | 5 | 0.0000000 | 16.557442 | 0.3393861 | 0.0000000 | 9.466947 | 1.1847657 | 10 | 8 | 4 |
| 48 | 3 | 7 | 0.0000000 | 11.304930 | 0.3364190 | 0.0000000 | 3.882443 | 1.8976861 | 10 | 8 | 4 |
| 49 | 4 | 6 | 0.0000000 | 6.932281 | 0.2767308 | 0.0000000 | 4.751341 | 1.0837230 | 10 | 8 | 4 |
| 50 | 5 | 4 | 0.0000000 | 7.437496 | 0.1859374 | 0.0000000 | 7.437496 | 0.8162890 | 10 | 8 | 4 |
| 51 | 1 | 13 | 0.1711051 | 9.688685 | 0.6289244 | 0.0403352 | 5.063579 | 2.2831273 | 11 | 4 | 5 |
| 52 | 2 | 7 | 0.0000000 | 11.322431 | 0.4471823 | 0.0000000 | 5.722932 | 1.9471475 | 11 | 4 | 5 |
| 53 | 3 | 11 | 0.1017516 | 14.993140 | 0.7521643 | 0.0000000 | 11.329369 | 2.7976261 | 11 | 4 | 5 |
| 54 | 4 | 6 | 0.0000000 | 7.155858 | 0.2681790 | 0.0000000 | 6.212797 | 0.9218882 | 11 | 4 | 5 |
| 55 | 5 | 8 | 0.0000000 | 9.299128 | 0.2772376 | 0.0000000 | 7.062117 | 1.6075429 | 11 | 4 | 5 |
| 56 | 1 | 9 | 0.0000000 | 14.361497 | 0.4451664 | 0.0000000 | 5.909446 | 2.0694053 | 12 | 5 | 5 |
| 57 | 2 | 8 | 0.0000000 | 10.902873 | 0.4167546 | 0.0000000 | 5.040122 | 1.6635569 | 12 | 5 | 5 |
| 58 | 3 | 8 | 0.0000000 | 13.200172 | 0.5846602 | 0.0000000 | 8.233837 | 1.9542346 | 12 | 5 | 5 |
| 59 | 4 | 5 | 0.0000000 | 7.220270 | 0.2624765 | 0.0000000 | 8.605687 | 0.8995180 | 12 | 5 | 5 |
| 60 | 5 | 5 | 0.0000000 | 7.764737 | 0.2111390 | 0.0000000 | 7.764737 | 0.9782088 | 12 | 5 | 5 |
| 61 | 1 | 4 | 0.0000000 | 16.756011 | 0.3497540 | 0.0000000 | 7.594901 | 1.6885375 | 13 | 6 | 5 |
| 62 | 2 | 7 | 0.0000000 | 13.951263 | 0.3748622 | 0.0000000 | 9.878982 | 1.8347756 | 13 | 6 | 5 |
| 63 | 3 | 11 | 0.0000000 | 10.921073 | 0.4054084 | 0.0000000 | 5.471170 | 2.1742571 | 13 | 6 | 5 |
| 64 | 4 | 6 | 0.0000000 | 6.932281 | 0.2783137 | 0.0000000 | 6.932281 | 1.0897496 | 13 | 6 | 5 |
| 65 | 5 | 4 | 0.0000000 | 8.753199 | 0.2040884 | 0.0000000 | 7.926292 | 0.9878955 | 13 | 6 | 5 |
| 66 | 1 | 5 | 0.0000000 | 14.928869 | 0.3185728 | 0.0000000 | 8.986369 | 1.5562112 | 14 | 7 | 5 |
| 67 | 2 | 5 | 0.0000000 | 16.557442 | 0.2877523 | 0.0000000 | 9.466947 | 1.2655765 | 14 | 7 | 5 |
| 68 | 3 | 8 | 0.0000000 | 10.066787 | 0.3296504 | 0.0000000 | 3.468666 | 2.0459750 | 14 | 7 | 5 |
| 69 | 4 | 6 | 0.0000000 | 6.932281 | 0.2791717 | 0.0000000 | 6.932281 | 1.2656769 | 14 | 7 | 5 |
| 70 | 5 | 4 | 0.0000000 | 7.539044 | 0.1602788 | 0.0000000 | 5.772494 | 0.5339783 | 14 | 7 | 5 |
| 71 | 1 | 3 | 0.0000000 | 20.737360 | 0.2420334 | 0.0000000 | 8.986369 | 1.2630193 | 15 | 8 | 5 |
| 72 | 2 | 5 | 0.0000000 | 16.557442 | 0.2715089 | 0.0000000 | 9.466947 | 1.1847657 | 15 | 8 | 5 |
| 73 | 3 | 7 | 0.0000000 | 11.304930 | 0.2691352 | 0.0000000 | 3.882443 | 1.8976861 | 15 | 8 | 5 |
| 74 | 4 | 6 | 0.0000000 | 6.932281 | 0.2213847 | 0.0000000 | 4.751341 | 1.0837230 | 15 | 8 | 5 |
| 75 | 5 | 4 | 0.0000000 | 7.437496 | 0.1487499 | 0.0000000 | 7.437496 | 0.8162890 | 15 | 8 | 5 |
Sensitivity analysis
sensitivity <- read.csv(
file.path("scenarios_resources", "r_sensitivity_results.csv")
)
kable(sensitivity) |> scroll_box(height = "400px")| X | replication | arrivals | mean_wait_time_doctor | mean_time_with_doctor | utilisation_doctor | mean_queue_length_doctor | mean_time_in_system | mean_patients_in_system | scenario | interarrival_time |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 1 | 8 | 0.0000000 | 10.626767 | 0.4811191 | 0.0000000 | 5.909446 | 1.5730708 | 1 | 4.0 |
| 2 | 2 | 14 | 1.3489715 | 16.474475 | 0.8415543 | 1.4037854 | 6.529895 | 4.0585764 | 1 | 4.0 |
| 3 | 3 | 8 | 3.3185568 | 12.576986 | 0.9214641 | 0.6425483 | 14.293369 | 2.4214309 | 1 | 4.0 |
| 4 | 4 | 9 | 7.0789803 | 8.229096 | 1.0000000 | 1.9960925 | 14.810041 | 3.2462020 | 1 | 4.0 |
| 5 | 5 | 11 | 11.8431217 | 8.866916 | 1.0000000 | 2.9270728 | 16.007073 | 4.2730406 | 1 | 4.0 |
| 6 | 1 | 11 | 4.4136200 | 10.968638 | 0.9305843 | 0.9874912 | 10.970224 | 3.2130228 | 2 | 4.5 |
| 7 | 2 | 10 | 4.2952348 | 11.628385 | 0.8801942 | 1.1333248 | 10.737537 | 3.5719202 | 2 | 4.5 |
| 8 | 3 | 8 | 6.3476572 | 13.246809 | 0.9553582 | 1.1892971 | 16.012763 | 3.1906477 | 2 | 4.5 |
| 9 | 4 | 4 | 1.6258165 | 9.999398 | 0.6295231 | 2.1705516 | 8.013161 | 0.7738957 | 2 | 4.5 |
| 10 | 5 | 7 | 1.4030508 | 7.662403 | 0.7949871 | 0.1776756 | 8.671934 | 1.8245624 | 2 | 4.5 |
| 11 | 1 | 9 | 1.1561321 | 12.600360 | 0.7545527 | 0.3739683 | 6.972457 | 2.4937655 | 3 | 5.0 |
| 12 | 2 | 10 | 0.2185845 | 15.462225 | 0.8137894 | 0.7190226 | 7.019465 | 3.0715488 | 3 | 5.0 |
| 13 | 3 | 8 | 1.6612341 | 11.704709 | 0.8144131 | 0.2671798 | 10.033307 | 1.7999155 | 3 | 5.0 |
| 14 | 4 | 4 | 1.3862671 | 8.268402 | 0.5633356 | 0.9223766 | 8.714268 | 0.8490077 | 3 | 5.0 |
| 15 | 5 | 7 | 1.1383003 | 8.211288 | 0.8027837 | 0.1753468 | 8.159880 | 1.5179933 | 3 | 5.0 |
| 16 | 1 | 8 | 0.0000000 | 13.183604 | 0.6555458 | 0.0736313 | 7.594901 | 2.0286140 | 4 | 5.5 |
| 17 | 2 | 6 | 0.0000000 | 14.204232 | 0.6476341 | 0.0000000 | 8.061193 | 1.7041228 | 4 | 5.5 |
| 18 | 3 | 9 | 1.0701721 | 10.484981 | 0.7546664 | 0.2840460 | 7.720035 | 2.5252492 | 4 | 5.5 |
| 19 | 4 | 8 | 1.6097542 | 12.349744 | 0.6931755 | 0.2616702 | 7.074266 | 2.0253240 | 4 | 5.5 |
| 20 | 5 | 7 | 0.6407407 | 8.647519 | 0.7528266 | 0.0000000 | 7.929976 | 1.3341535 | 4 | 5.5 |
| 21 | 1 | 4 | 0.0000000 | 16.756011 | 0.5829234 | 0.0000000 | 7.594901 | 1.6885375 | 5 | 6.0 |
| 22 | 2 | 7 | 0.0000000 | 13.951263 | 0.6247704 | 0.0000000 | 9.878982 | 1.8347756 | 5 | 6.0 |
| 23 | 3 | 11 | 0.0000000 | 10.642901 | 0.6696266 | 0.0000000 | 5.471170 | 2.1742571 | 5 | 6.0 |
| 24 | 4 | 5 | 2.1255795 | 7.962361 | 0.4490071 | 0.2915664 | 10.087940 | 1.3321312 | 5 | 6.0 |
| 25 | 5 | 6 | 0.1698502 | 7.931855 | 0.6664404 | 0.0000000 | 7.985399 | 0.9107067 | 5 | 6.0 |
| 26 | 1 | 5 | 0.0000000 | 14.928869 | 0.5671720 | 0.0000000 | 7.601250 | 1.5875062 | 6 | 6.5 |
| 27 | 2 | 6 | 0.0000000 | 15.719055 | 0.5351976 | 0.0000000 | 12.100828 | 1.5937329 | 6 | 6.5 |
| 28 | 3 | 8 | 0.0000000 | 10.066787 | 0.5907619 | 0.0000000 | 5.471170 | 2.0509018 | 6 | 6.5 |
| 29 | 4 | 5 | 1.5138841 | 7.962361 | 0.4497221 | 0.3399041 | 9.476245 | 1.3465966 | 6 | 6.5 |
| 30 | 5 | 5 | 0.0046527 | 13.128797 | 0.6221594 | 0.0000000 | 13.133450 | 1.7319608 | 6 | 6.5 |
| 31 | 1 | 5 | 0.0000000 | 14.928869 | 0.5309547 | 0.0000000 | 8.986369 | 1.5562112 | 7 | 7.0 |
| 32 | 2 | 5 | 0.0000000 | 16.557442 | 0.4795871 | 0.0000000 | 9.466947 | 1.2655765 | 7 | 7.0 |
| 33 | 3 | 8 | 0.0000000 | 10.066787 | 0.5494173 | 0.0000000 | 3.468666 | 2.0459750 | 7 | 7.0 |
| 34 | 4 | 5 | 0.9021887 | 7.962361 | 0.4504371 | 0.3962059 | 8.864549 | 1.3634454 | 7 | 7.0 |
| 35 | 5 | 4 | 0.0000000 | 8.753199 | 0.3340787 | 0.0000000 | 7.926292 | 0.8053229 | 7 | 7.0 |
Example tables
This function finds the average results from each experiment.
It uses the summary_stats() function introduced on the Replications page.
summary_stats()
def summary_stats(data):
"""
Calculate mean, standard deviation and 95% confidence interval (CI).
Parameters
----------
data : pd.Series
Data to use in calculation.
Returns
-------
tuple
(mean, standard deviation, CI lower, CI upper).
"""
# Remove any NaN from the series
data = data.dropna()
# Find number of observations
count = len(data)
# If there are no observations, then set all to NaN
if count == 0:
mean, std_dev, ci_lower, ci_upper = np.nan, np.nan, np.nan, np.nan
# If there is only one or two observations, can do mean but not others
elif count < 3:
mean = data.mean()
std_dev, ci_lower, ci_upper = np.nan, np.nan, np.nan
# With more than one observation, can calculate all...
else:
mean = data.mean()
std_dev = data.std()
# Special case for CI if variance is 0
if np.var(data) == 0:
ci_lower, ci_upper = mean, mean
else:
# Calculation of CI uses t-distribution, which is suitable for
# smaller sample sizes (n<30)
ci_lower, ci_upper = st.t.interval(
confidence=0.95,
df=count-1,
loc=mean,
scale=st.sem(data))
return mean, std_dev, ci_lower, ci_upperdef summarise_scenarios(results, groups, result_vars, path_prefix=None):
"""
Find the average results from each scenario for multiple metrics.
Parameters
----------
results : pd.DataFrame
Run-level results from every scenario.
groups : list
List of columns to group by (provided as strings).
result_vars : list
List of performance measures to get results on (provided as strings).
path_prefix : str, optional
Path prefix to save tables to. Each metric will be saved as
{path_prefix}_{metric}.csv
Returns
-------
summary_tables : dict
Dictionary with metric names as keys and summary DataFrames as values.
"""
summary_tables = {}
for result_var in result_vars:
summary_table = (
results.groupby(groups)[result_var]
.apply(summary_stats)
.apply(pd.Series)
.reset_index()
)
summary_table.columns = (
list(summary_table.columns[:-4]) +
["mean", "std_dev", "ci_lower", "ci_upper"]
)
# Add column to identify which metric this is
summary_table["metric"] = result_var
summary_tables[result_var] = summary_table
# Save if path provided
if path_prefix:
output_path = f"{path_prefix}_{result_var}.csv"
summary_table.to_csv(output_path, index=False)
return summary_tablesScenario analysis
Here and throughout page, we will save results to a folder called tables_figures_resources, and then a subfolder (e.g., python_scenario/). You can rename this path (e.g. to os.path.join("results", "scenario"))), or otherwise, create the listed directory structure (here, tables_figures_resources/python_scenario/).
result_variables = [
"mean_wait_time",
"mean_utilisation_tw",
"mean_queue_length",
"mean_time_in_system",
"mean_patients_in_system"
]scenario_tables = summarise_scenarios(
results=scenario,
groups=["scenario", "interarrival_time", "number_of_doctors"],
result_vars=result_variables,
path_prefix=os.path.join("tables_figures_resources", "python_scenario")
)Click the name of each metric below to view the relevant dataframe:
HTML(to_html_datatable(scenario_tables["mean_wait_time"]))| Loading ITables v2.5.2 from the internet... (need help?) |
HTML(to_html_datatable(scenario_tables["mean_utilisation_tw"]))| Loading ITables v2.5.2 from the internet... (need help?) |
HTML(to_html_datatable(scenario_tables["mean_queue_length"]))| Loading ITables v2.5.2 from the internet... (need help?) |
HTML(to_html_datatable(scenario_tables["mean_time_in_system"]))| Loading ITables v2.5.2 from the internet... (need help?) |
HTML(to_html_datatable(scenario_tables["mean_patients_in_system"]))| Loading ITables v2.5.2 from the internet... (need help?) |
Sensitivity analysis
sensitivity_tables = summarise_scenarios(
results=sensitivity,
groups=["scenario", "interarrival_time"],
result_vars=result_variables,
path_prefix=os.path.join("tables_figures_resources", "python_sensitivity")
)Click the name of each metric below to view the relevant dataframe:
HTML(to_html_datatable(sensitivity_tables["mean_wait_time"]))| Loading ITables v2.5.2 from the internet... (need help?) |
HTML(to_html_datatable(sensitivity_tables["mean_utilisation_tw"]))| Loading ITables v2.5.2 from the internet... (need help?) |
HTML(to_html_datatable(sensitivity_tables["mean_queue_length"]))| Loading ITables v2.5.2 from the internet... (need help?) |
HTML(to_html_datatable(sensitivity_tables["mean_time_in_system"]))| Loading ITables v2.5.2 from the internet... (need help?) |
HTML(to_html_datatable(sensitivity_tables["mean_patients_in_system"]))| Loading ITables v2.5.2 from the internet... (need help?) |
#' Summarise multiple performance measures at once
#'
#' @param results Dataframe with results from each replication of scenarios.
#' @param groups Vector of grouping columns (as strings).
#' @param result_vars Vector of performance measures to summarise (as strings).
#' @param path_prefix Optional path prefix to save each table to.
#'
#' @return A named list of summary data frames.
summarise_scenarios <- function(
results, groups, result_vars, path_prefix = NULL
) {
summary_tables <- list()
for (result_var in result_vars) {
summary_table <- results |>
group_by(across(all_of(groups))) |>
summarise(
mean = mean(.data[[result_var]], na.rm = TRUE),
std_dev = sd(.data[[result_var]], na.rm = TRUE),
ci_lower = t.test(.data[[result_var]])$conf.int[1L],
ci_upper = t.test(.data[[result_var]])$conf.int[2L],
.groups = "drop"
) |>
mutate(metric = result_var)
# Save table if prefix specified
if (!is.null(path_prefix)) {
output_path <- paste0(path_prefix, "_", result_var, ".csv")
write.csv(summary_table, output_path, row.names = FALSE)
}
# Store in list
summary_tables[[result_var]] <- summary_table
}
return(summary_tables)
}Scenario analysis
result_variables <- c(
"mean_wait_time_doctor",
"utilisation_doctor",
"mean_queue_length_doctor",
"mean_time_in_system",
"mean_patients_in_system"
)scenario_tables <- summarise_scenarios(
results = scenario,
groups = c("scenario", "interarrival_time", "number_of_doctors"),
result_vars = result_variables,
path_prefix = file.path("tables_figures_resources", "r_scenario")
)Click the name of each metric below to view the relevant table:
kable(scenario_tables[["mean_wait_time_doctor"]])| scenario | interarrival_time | number_of_doctors | mean | std_dev | ci_lower | ci_upper | metric |
|---|---|---|---|---|---|---|---|
| 1 | 4 | 3 | 4.7179260 | 4.7934827 | -1.2339688 | 10.6698209 | mean_wait_time_doctor |
| 2 | 5 | 3 | 1.1121036 | 0.5426120 | 0.4383618 | 1.7858454 | mean_wait_time_doctor |
| 3 | 6 | 3 | 0.4590859 | 0.9344969 | -0.7012452 | 1.6194171 | mean_wait_time_doctor |
| 4 | 7 | 3 | 0.1804377 | 0.4034711 | -0.3205377 | 0.6814132 | mean_wait_time_doctor |
| 5 | 8 | 3 | 0.0000000 | 0.0000000 | NaN | NaN | mean_wait_time_doctor |
| 6 | 4 | 4 | 0.1230492 | 0.1691037 | -0.0869207 | 0.3330191 | mean_wait_time_doctor |
| 7 | 5 | 4 | 0.0200439 | 0.0448196 | -0.0356069 | 0.0756948 | mean_wait_time_doctor |
| 8 | 6 | 4 | 0.0000000 | 0.0000000 | NaN | NaN | mean_wait_time_doctor |
| 9 | 7 | 4 | 0.0000000 | 0.0000000 | NaN | NaN | mean_wait_time_doctor |
| 10 | 8 | 4 | 0.0000000 | 0.0000000 | NaN | NaN | mean_wait_time_doctor |
| 11 | 4 | 5 | 0.0545714 | 0.0786451 | -0.0430794 | 0.1522221 | mean_wait_time_doctor |
| 12 | 5 | 5 | 0.0000000 | 0.0000000 | NaN | NaN | mean_wait_time_doctor |
| 13 | 6 | 5 | 0.0000000 | 0.0000000 | NaN | NaN | mean_wait_time_doctor |
| 14 | 7 | 5 | 0.0000000 | 0.0000000 | NaN | NaN | mean_wait_time_doctor |
| 15 | 8 | 5 | 0.0000000 | 0.0000000 | NaN | NaN | mean_wait_time_doctor |
kable(scenario_tables[["utilisation_doctor"]])| scenario | interarrival_time | number_of_doctors | mean | std_dev | ci_lower | ci_upper | metric |
|---|---|---|---|---|---|---|---|
| 1 | 4 | 3 | 0.8488275 | 0.2157804 | 0.5809008 | 1.1167542 | utilisation_doctor |
| 2 | 5 | 3 | 0.7497749 | 0.1070844 | 0.6168120 | 0.8827378 | utilisation_doctor |
| 3 | 6 | 3 | 0.5985536 | 0.0907686 | 0.4858495 | 0.7112577 | utilisation_doctor |
| 4 | 7 | 3 | 0.4688950 | 0.0850966 | 0.3632336 | 0.5745563 | utilisation_doctor |
| 5 | 8 | 3 | 0.3927330 | 0.0668498 | 0.3097280 | 0.4757381 | utilisation_doctor |
| 6 | 4 | 4 | 0.5597782 | 0.2209634 | 0.2854159 | 0.8341405 | utilisation_doctor |
| 7 | 5 | 4 | 0.4790470 | 0.1855439 | 0.2486638 | 0.7094302 | utilisation_doctor |
| 8 | 6 | 4 | 0.4031067 | 0.1014199 | 0.2771773 | 0.5290361 | utilisation_doctor |
| 9 | 7 | 4 | 0.3438565 | 0.0843777 | 0.2390877 | 0.4486253 | utilisation_doctor |
| 10 | 8 | 4 | 0.2882030 | 0.0627295 | 0.2103141 | 0.3660920 | utilisation_doctor |
| 11 | 4 | 5 | 0.4747375 | 0.2139925 | 0.2090308 | 0.7404443 | utilisation_doctor |
| 12 | 5 | 5 | 0.3840393 | 0.1497728 | 0.1980718 | 0.5700069 | utilisation_doctor |
| 13 | 6 | 5 | 0.3224854 | 0.0811359 | 0.2217418 | 0.4232289 | utilisation_doctor |
| 14 | 7 | 5 | 0.2750852 | 0.0675022 | 0.1912702 | 0.3589002 | utilisation_doctor |
| 15 | 8 | 5 | 0.2305624 | 0.0501836 | 0.1682513 | 0.2928736 | utilisation_doctor |
kable(scenario_tables[["mean_queue_length_doctor"]])| scenario | interarrival_time | number_of_doctors | mean | std_dev | ci_lower | ci_upper | metric |
|---|---|---|---|---|---|---|---|
| 1 | 4 | 3 | 1.3938998 | 1.1424578 | -0.0246489 | 2.8124485 | mean_queue_length_doctor |
| 2 | 5 | 3 | 0.4915788 | 0.3168230 | 0.0981911 | 0.8849665 | mean_queue_length_doctor |
| 3 | 6 | 3 | 0.0583133 | 0.1303925 | -0.1035904 | 0.2202169 | mean_queue_length_doctor |
| 4 | 7 | 3 | 0.0792412 | 0.1771887 | -0.1407676 | 0.2992500 | mean_queue_length_doctor |
| 5 | 8 | 3 | 0.0000000 | 0.0000000 | NaN | NaN | mean_queue_length_doctor |
| 6 | 4 | 4 | 0.0000000 | 0.0000000 | NaN | NaN | mean_queue_length_doctor |
| 7 | 5 | 4 | 0.0000000 | 0.0000000 | NaN | NaN | mean_queue_length_doctor |
| 8 | 6 | 4 | 0.0000000 | 0.0000000 | NaN | NaN | mean_queue_length_doctor |
| 9 | 7 | 4 | 0.0000000 | 0.0000000 | NaN | NaN | mean_queue_length_doctor |
| 10 | 8 | 4 | 0.0000000 | 0.0000000 | NaN | NaN | mean_queue_length_doctor |
| 11 | 4 | 5 | 0.0080670 | 0.0180384 | -0.0143306 | 0.0304647 | mean_queue_length_doctor |
| 12 | 5 | 5 | 0.0000000 | 0.0000000 | NaN | NaN | mean_queue_length_doctor |
| 13 | 6 | 5 | 0.0000000 | 0.0000000 | NaN | NaN | mean_queue_length_doctor |
| 14 | 7 | 5 | 0.0000000 | 0.0000000 | NaN | NaN | mean_queue_length_doctor |
| 15 | 8 | 5 | 0.0000000 | 0.0000000 | NaN | NaN | mean_queue_length_doctor |
kable(scenario_tables[["mean_time_in_system"]])| scenario | interarrival_time | number_of_doctors | mean | std_dev | ci_lower | ci_upper | metric |
|---|---|---|---|---|---|---|---|
| 1 | 4 | 3 | 11.509965 | 4.874134 | 5.457928 | 17.562002 | mean_time_in_system |
| 2 | 5 | 3 | 8.179875 | 1.277262 | 6.593945 | 9.765806 | mean_time_in_system |
| 3 | 6 | 3 | 8.203678 | 1.886925 | 5.860752 | 10.546605 | mean_time_in_system |
| 4 | 7 | 3 | 7.742565 | 2.453536 | 4.696097 | 10.789032 | mean_time_in_system |
| 5 | 8 | 3 | 7.002678 | 2.533037 | 3.857498 | 10.147859 | mean_time_in_system |
| 6 | 4 | 4 | 7.028795 | 2.374920 | 4.079942 | 9.977648 | mean_time_in_system |
| 7 | 5 | 4 | 7.110766 | 1.553584 | 5.181737 | 9.039794 | mean_time_in_system |
| 8 | 6 | 4 | 7.560725 | 1.602366 | 5.571126 | 9.550325 | mean_time_in_system |
| 9 | 7 | 4 | 6.925351 | 2.448739 | 3.884840 | 9.965863 | mean_time_in_system |
| 10 | 8 | 4 | 6.904919 | 2.497665 | 3.803659 | 10.006180 | mean_time_in_system |
| 11 | 4 | 5 | 7.078159 | 2.485813 | 3.991614 | 10.164704 | mean_time_in_system |
| 12 | 5 | 5 | 7.110766 | 1.553584 | 5.181737 | 9.039794 | mean_time_in_system |
| 13 | 6 | 5 | 7.560725 | 1.602366 | 5.571126 | 9.550325 | mean_time_in_system |
| 14 | 7 | 5 | 6.925351 | 2.448739 | 3.884840 | 9.965863 | mean_time_in_system |
| 15 | 8 | 5 | 6.904919 | 2.497665 | 3.803659 | 10.006180 | mean_time_in_system |
kable(scenario_tables[["mean_patients_in_system"]])| scenario | interarrival_time | number_of_doctors | mean | std_dev | ci_lower | ci_upper | metric |
|---|---|---|---|---|---|---|---|
| 1 | 4 | 3 | 3.114464 | 1.1299547 | 1.7114401 | 4.517488 | mean_patients_in_system |
| 2 | 5 | 3 | 1.946446 | 0.8623846 | 0.8756543 | 3.017238 | mean_patients_in_system |
| 3 | 6 | 3 | 1.588082 | 0.4844839 | 0.9865155 | 2.189648 | mean_patients_in_system |
| 4 | 7 | 3 | 1.407306 | 0.4512387 | 0.8470193 | 1.967593 | mean_patients_in_system |
| 5 | 8 | 3 | 1.205324 | 0.4656828 | 0.6271028 | 1.783546 | mean_patients_in_system |
| 6 | 4 | 4 | 1.779714 | 0.5515137 | 1.0949191 | 2.464509 | mean_patients_in_system |
| 7 | 5 | 4 | 1.512985 | 0.5452761 | 0.8359350 | 2.190034 | mean_patients_in_system |
| 8 | 6 | 4 | 1.555043 | 0.5043936 | 0.9287557 | 2.181330 | mean_patients_in_system |
| 9 | 7 | 4 | 1.333484 | 0.5490149 | 0.6517916 | 2.015176 | mean_patients_in_system |
| 10 | 8 | 4 | 1.249097 | 0.3998974 | 0.7525585 | 1.745635 | mean_patients_in_system |
| 11 | 4 | 5 | 1.911466 | 0.7064557 | 1.0342858 | 2.788647 | mean_patients_in_system |
| 12 | 5 | 5 | 1.512985 | 0.5452761 | 0.8359350 | 2.190034 | mean_patients_in_system |
| 13 | 6 | 5 | 1.555043 | 0.5043936 | 0.9287557 | 2.181330 | mean_patients_in_system |
| 14 | 7 | 5 | 1.333484 | 0.5490149 | 0.6517916 | 2.015176 | mean_patients_in_system |
| 15 | 8 | 5 | 1.249097 | 0.3998974 | 0.7525585 | 1.745635 | mean_patients_in_system |
Sensitivity analysis
sensitivity_tables <- summarise_scenarios(
results = sensitivity,
groups = c("scenario", "interarrival_time"),
result_vars = result_variables,
path_prefix = file.path("tables_figures_resources", "r_sensitivity")
)Click the name of each metric below to view the relevant table:
kable(sensitivity_tables[["mean_wait_time_doctor"]])| scenario | interarrival_time | mean | std_dev | ci_lower | ci_upper | metric |
|---|---|---|---|---|---|---|
| 1 | 4.0 | 4.7179260 | 4.7934827 | -1.2339688 | 10.6698209 | mean_wait_time_doctor |
| 2 | 4.5 | 3.6170759 | 2.0867126 | 1.0260800 | 6.2080718 | mean_wait_time_doctor |
| 3 | 5.0 | 1.1121036 | 0.5426120 | 0.4383618 | 1.7858454 | mean_wait_time_doctor |
| 4 | 5.5 | 0.6641334 | 0.6967352 | -0.2009776 | 1.5292444 | mean_wait_time_doctor |
| 5 | 6.0 | 0.4590859 | 0.9344969 | -0.7012452 | 1.6194171 | mean_wait_time_doctor |
| 6 | 6.5 | 0.3037073 | 0.6765124 | -0.5362937 | 1.1437084 | mean_wait_time_doctor |
| 7 | 7.0 | 0.1804377 | 0.4034711 | -0.3205377 | 0.6814132 | mean_wait_time_doctor |
kable(sensitivity_tables[["utilisation_doctor"]])| scenario | interarrival_time | mean | std_dev | ci_lower | ci_upper | metric |
|---|---|---|---|---|---|---|
| 1 | 4.0 | 0.8488275 | 0.2157804 | 0.5809008 | 1.1167542 | utilisation_doctor |
| 2 | 4.5 | 0.8381294 | 0.1317552 | 0.6745337 | 1.0017251 | utilisation_doctor |
| 3 | 5.0 | 0.7497749 | 0.1070844 | 0.6168120 | 0.8827378 | utilisation_doctor |
| 4 | 5.5 | 0.7007697 | 0.0513348 | 0.6370291 | 0.7645102 | utilisation_doctor |
| 5 | 6.0 | 0.5985536 | 0.0907686 | 0.4858495 | 0.7112577 | utilisation_doctor |
| 6 | 6.5 | 0.5530026 | 0.0659414 | 0.4711256 | 0.6348797 | utilisation_doctor |
| 7 | 7.0 | 0.4688950 | 0.0850966 | 0.3632336 | 0.5745563 | utilisation_doctor |
kable(sensitivity_tables[["mean_queue_length_doctor"]])| scenario | interarrival_time | mean | std_dev | ci_lower | ci_upper | metric |
|---|---|---|---|---|---|---|
| 1 | 4.0 | 1.3938998 | 1.1424578 | -0.0246489 | 2.8124485 | mean_queue_length_doctor |
| 2 | 4.5 | 1.1316681 | 0.7094877 | 0.2507227 | 2.0126135 | mean_queue_length_doctor |
| 3 | 5.0 | 0.4915788 | 0.3168230 | 0.0981911 | 0.8849665 | mean_queue_length_doctor |
| 4 | 5.5 | 0.1238695 | 0.1395141 | -0.0493601 | 0.2970991 | mean_queue_length_doctor |
| 5 | 6.0 | 0.0583133 | 0.1303925 | -0.1035904 | 0.2202169 | mean_queue_length_doctor |
| 6 | 6.5 | 0.0679808 | 0.1520097 | -0.1207642 | 0.2567258 | mean_queue_length_doctor |
| 7 | 7.0 | 0.0792412 | 0.1771887 | -0.1407676 | 0.2992500 | mean_queue_length_doctor |
kable(sensitivity_tables[["mean_time_in_system"]])| scenario | interarrival_time | mean | std_dev | ci_lower | ci_upper | metric |
|---|---|---|---|---|---|---|
| 1 | 4.0 | 11.509965 | 4.8741341 | 5.457928 | 17.562002 | mean_time_in_system |
| 2 | 4.5 | 10.881124 | 3.1411782 | 6.980836 | 14.781412 | mean_time_in_system |
| 3 | 5.0 | 8.179875 | 1.2772625 | 6.593945 | 9.765806 | mean_time_in_system |
| 4 | 5.5 | 7.676074 | 0.3819287 | 7.201847 | 8.150301 | mean_time_in_system |
| 5 | 6.0 | 8.203678 | 1.8869246 | 5.860752 | 10.546605 | mean_time_in_system |
| 6 | 6.5 | 9.556589 | 3.1538682 | 5.640544 | 13.472633 | mean_time_in_system |
| 7 | 7.0 | 7.742565 | 2.4535360 | 4.696097 | 10.789032 | mean_time_in_system |
kable(sensitivity_tables[["mean_patients_in_system"]])| scenario | interarrival_time | mean | std_dev | ci_lower | ci_upper | metric |
|---|---|---|---|---|---|---|
| 1 | 4.0 | 3.114464 | 1.1299547 | 1.7114401 | 4.517488 | mean_patients_in_system |
| 2 | 4.5 | 2.514810 | 1.1799326 | 1.0497299 | 3.979890 | mean_patients_in_system |
| 3 | 5.0 | 1.946446 | 0.8623846 | 0.8756543 | 3.017238 | mean_patients_in_system |
| 4 | 5.5 | 1.923493 | 0.4412978 | 1.3755491 | 2.471436 | mean_patients_in_system |
| 5 | 6.0 | 1.588082 | 0.4844839 | 0.9865155 | 2.189648 | mean_patients_in_system |
| 6 | 6.5 | 1.662140 | 0.2577926 | 1.3420479 | 1.982231 | mean_patients_in_system |
| 7 | 7.0 | 1.407306 | 0.4512387 | 0.8470193 | 1.967593 | mean_patients_in_system |
Example figures
These functions are used to plot the results from the scenarios and sensitivity analysis.
We have used plotly.express for our plots.
def pale_colour(colour, alpha=0.2):
"""
Make pale version of a colour.
Parameters
----------
colour : str or tuple
The colour to pale, as a Plotly-compatible name ('blue', 'red'), hex
string (e.g., "#1f77b4"), or RGB tuple (e.g., (0.1, 0.3, 0.8)).
Accepts any format readable by matplotlib.colors.to_rgb.
alpha : float, optional
Opacity for the pale colour (default = 0.2). Should be between 0 and 1.
Returns
-------
str
Colour in 'rgba(r, g, b, a)' string format
(e.g., "rgba(31,119,180,0.2)").
Acknowledgements
----------------
This function was generated by Perplexity.
"""
# Convert any accepted colour format to an RGB tuple
rgb = mcolors.to_rgb(colour)
# Scale values to 0-255 for plotly compatability
r, g, b = [int(x * 255) for x in rgb]
# Return as RGB tuple with added alpha parameter (which makes it paler)
return f"rgba({r},{g},{b},{alpha})"def plot_metrics(
summary_tables, colour_var=None, name_mappings=None, path_prefix=None
):
"""
Plot results from different model scenarios.
Parameters
----------
summary_tables : dict
Dictionary with metric names as keys and summary DataFrames as values.
colour_var : str, optional
Name of variable to colour lines with.
name_mappings : dict, optional
Dictionary mapping column names to labels. If not provided,
function will default to variable names.
path : str
Path to save figure to.
Returns
-------
figs : dict
Dictionary with metric names as keys and plotly figures as values.
"""
figs = {}
for metric_name, summary_table in summary_tables.items():
# Initialise figure
fig = go.Figure()
# Handling color mappings (optional)
if colour_var:
unique_groups = summary_table[colour_var].unique()
colours = px.colors.qualitative.Plotly
colour_map = {
group: colours[i % len(colours)]
for i, group in enumerate(unique_groups)
}
else:
colour_map = {None: "blue"}
# Plot each scenario line and its shaded confidence interval
if colour_var:
groups = summary_table.groupby(colour_var)
else:
groups = [(None, summary_table)]
for group_name, group_data in groups:
x = group_data["interarrival_time"].values
mean = group_data["mean"].values
ci_upper = group_data["ci_upper"].values
ci_lower = group_data["ci_lower"].values
main_colour = colour_map[group_name]
shade_colour = pale_colour(main_colour, alpha=0.2)
# Filled confidence interval region
fig.add_trace(go.Scatter(
x=np.concatenate([x, x[::-1]]),
y=np.concatenate([ci_upper, ci_lower[::-1]]),
fill="toself",
fillcolor=shade_colour,
line={"color": "rgba(255,255,255,0)"},
showlegend=False,
name="95% CI",
hoverinfo="name+y"
))
# Mean line
fig.add_trace(go.Scatter(
x=x,
y=mean,
mode="lines",
line={"color": main_colour, "width": 2},
name=group_name,
hoverinfo="y"
))
# Set labels and layout
fig.update_layout(
xaxis_title=(
name_mappings.get("interarrival_time", "interarrival_time")
if name_mappings else "interarrival_time"
),
yaxis_title=(
name_mappings.get(metric_name, metric_name)
if name_mappings else metric_name
),
legend_title_text=(
name_mappings.get(colour_var, colour_var)
if colour_var and name_mappings
else (colour_var if colour_var else None)
),
template="plotly_white"
)
if not colour_var:
fig.update_layout(showlegend=False)
figs[metric_name] = fig
# Save if path provided
if path_prefix:
output_path = f"{path_prefix}_{metric_name}.png"
fig.write_image(output_path)
return figsScenario analysis
name_mappings = {
"interarrival_time": "Patient inter-arrival time",
"number_of_doctors": "Number of doctors",
"mean_wait_time": "Mean wait time for the doctor",
"mean_utilisation_tw": "Mean utilisation",
"mean_queue_length": "Mean queue length",
"mean_time_in_system": "Mean patient time in the system",
"mean_patients_in_system": "Mean number of patients in the system"
}scenario_plots = plot_metrics(
summary_tables=scenario_tables,
colour_var="number_of_doctors",
name_mappings=name_mappings,
path_prefix=os.path.join("tables_figures_resources", "python_scenario")
)Click the name of each metric below to view the relevant plot:
scenario_plots["mean_wait_time"]scenario_plots["mean_utilisation_tw"]scenario_plots["mean_queue_length"]scenario_plots["mean_time_in_system"]scenario_plots["mean_patients_in_system"]This function is used to plot the results from the scenarios and sensitivity analysis.
We have used ggplot2 for our plots.
#' Plot multiple performance measures at once
#'
#' @param summary_tables Named list of summary tables, one per metric
#' (like output from summarise_scenarios()).
#' @param x_var Name of variable to plot on x axis.
#' @param colour_var Name of variable to colour lines with (can be NULL).
#' @param name_mappings Optional named list for prettier axis/legend labels.
#' @param path_prefix Optional path prefix to save figures.
#' @return Named list of ggplot objects
plot_metrics <- function(
summary_tables, x_var, colour_var = NULL, name_mappings = NULL,
path_prefix = NULL
) {
# List to store plots for each metric
plot_list <- list()
for (metric_name in names(summary_tables)) {
# Extract relevant results table
summary_table <- summary_tables[[metric_name]]
# Helper to map a variable to display name if available in `name_mappings`.
# Just uses variable name if no mapping is found.
get_name <- function(var) {
if (!is.null(var) && !is.null(name_mappings[[var]])) {
name_mappings[[var]]
} else {
var
}
}
xaxis_title <- get_name(x_var)
yaxis_title <- get_name(metric_name)
legend_title <- get_name(colour_var)
# Create plot, with or without grouping colour variable
if (!is.null(colour_var)) {
summary_table[[colour_var]] <- as.factor(summary_table[[colour_var]])
p <- ggplot(
summary_table,
aes_string(
x = x_var,
y = "mean",
group = colour_var,
color = colour_var,
fill = colour_var
)
) +
geom_line() +
geom_ribbon(
aes_string(
ymin = "ci_lower",
ymax = "ci_upper"
),
alpha = 0.1
) +
labs(
x = xaxis_title,
y = yaxis_title,
color = legend_title,
fill = legend_title
) +
theme_minimal()
} else {
p <- ggplot(
summary_table,
aes_string(
x = x_var,
y = "mean"
)
) +
geom_line() +
geom_ribbon(
aes_string(
ymin = "ci_lower",
ymax = "ci_upper"
),
alpha = 0.1,
show.legend = FALSE
) +
labs(
x = xaxis_title,
y = yaxis_title
) +
theme_minimal()
}
# Save plot if prefix supplied
if (!is.null(path_prefix)) {
output_path <- paste0(path_prefix, "_", metric_name, ".png")
ggsave(filename = output_path, plot = p, width = 6.5, height = 4L,
bg = "white")
}
plot_list[[metric_name]] <- p
}
return(plot_list)
}Scenario analysis
name_mappings <- list(
interarrival_time = "Patient inter-arrival time",
number_of_doctors = "Number of doctors",
mean_wait_time_doctor = "Mean wait time for the doctor",
utilisation_doctor = "Mean utilisation",
mean_queue_length_doctor = "Mean queue length",
mean_time_in_system = "Mean patient time in the system",
mean_patients_in_system = "Mean number of patients in the system"
)scenario_plots <- plot_metrics(
summary_tables = scenario_tables,
x_var = "interarrival_time",
colour_var = "number_of_doctors",
name_mappings = name_mappings,
path_prefix = file.path("tables_figures_resources", "r_scenario")
)Warning: `aes_string()` was deprecated in ggplot2 3.0.0.
ℹ Please use tidy evaluation idioms with `aes()`.
ℹ See also `vignette("ggplot2-in-packages")` for more information.Warning: Removed 8 rows containing missing values or values outside the scale range
(`geom_ribbon()`).Warning: Removed 10 rows containing missing values or values outside the scale range
(`geom_ribbon()`).Click the name of each metric below to view the relevant plot:
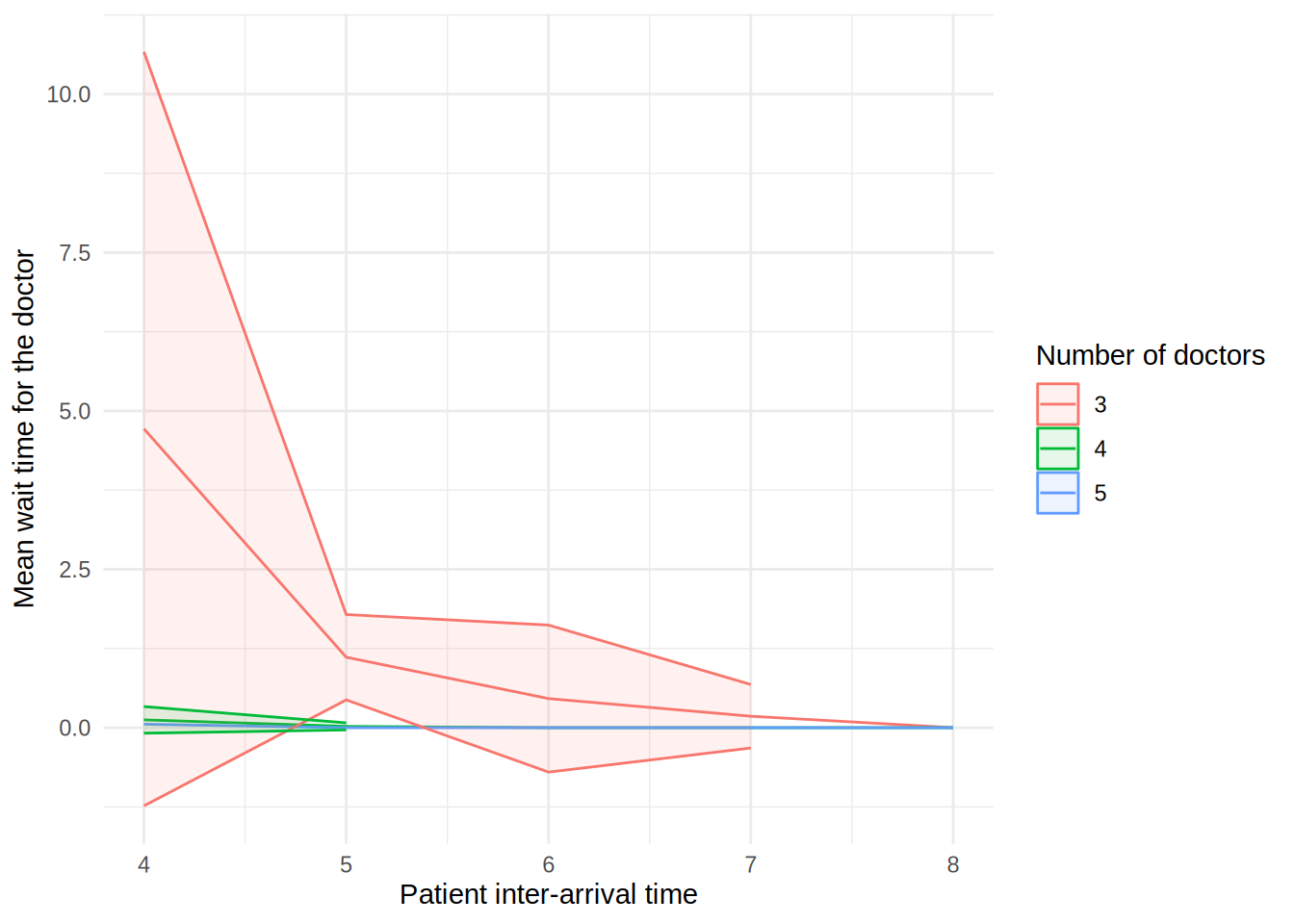
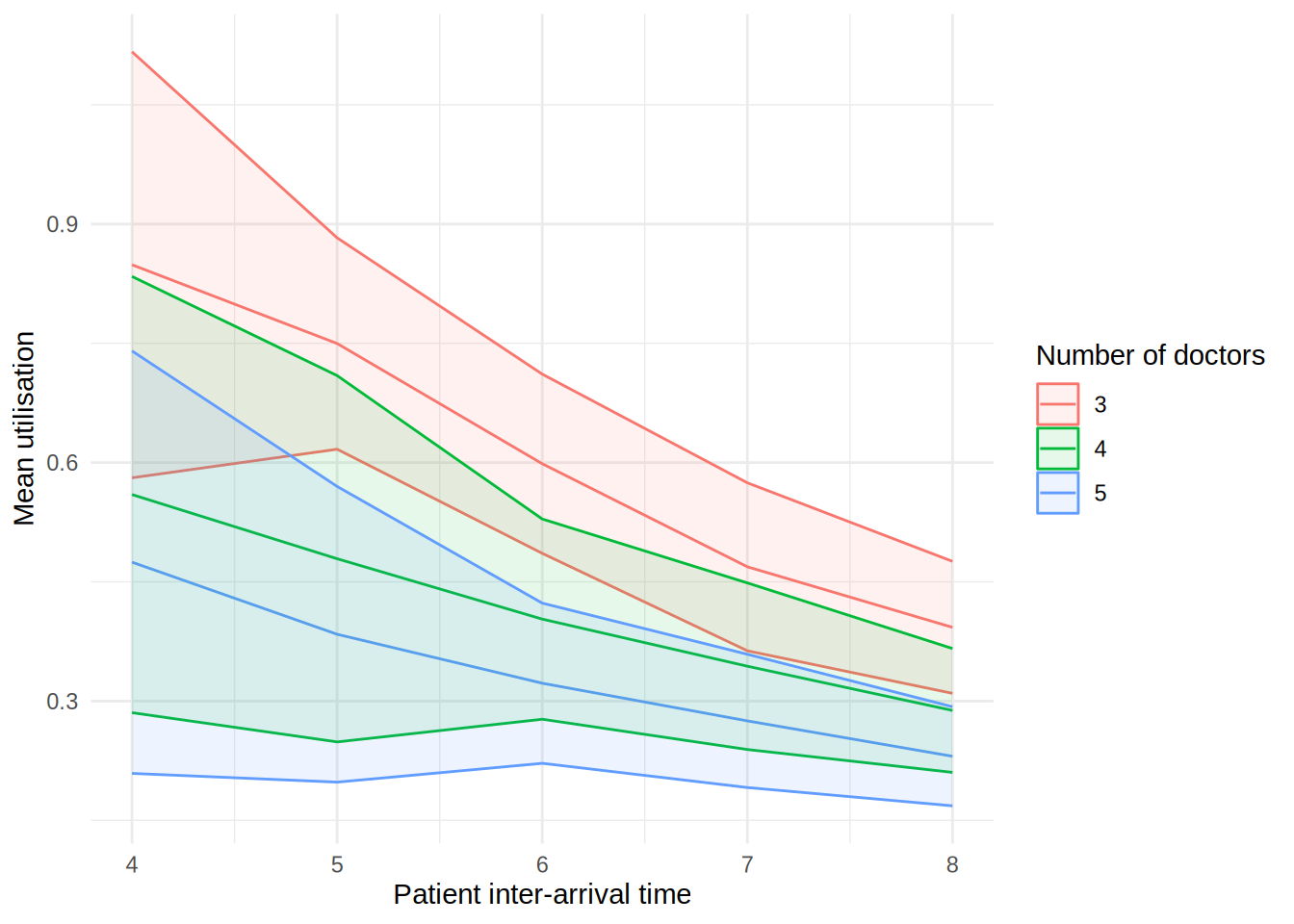
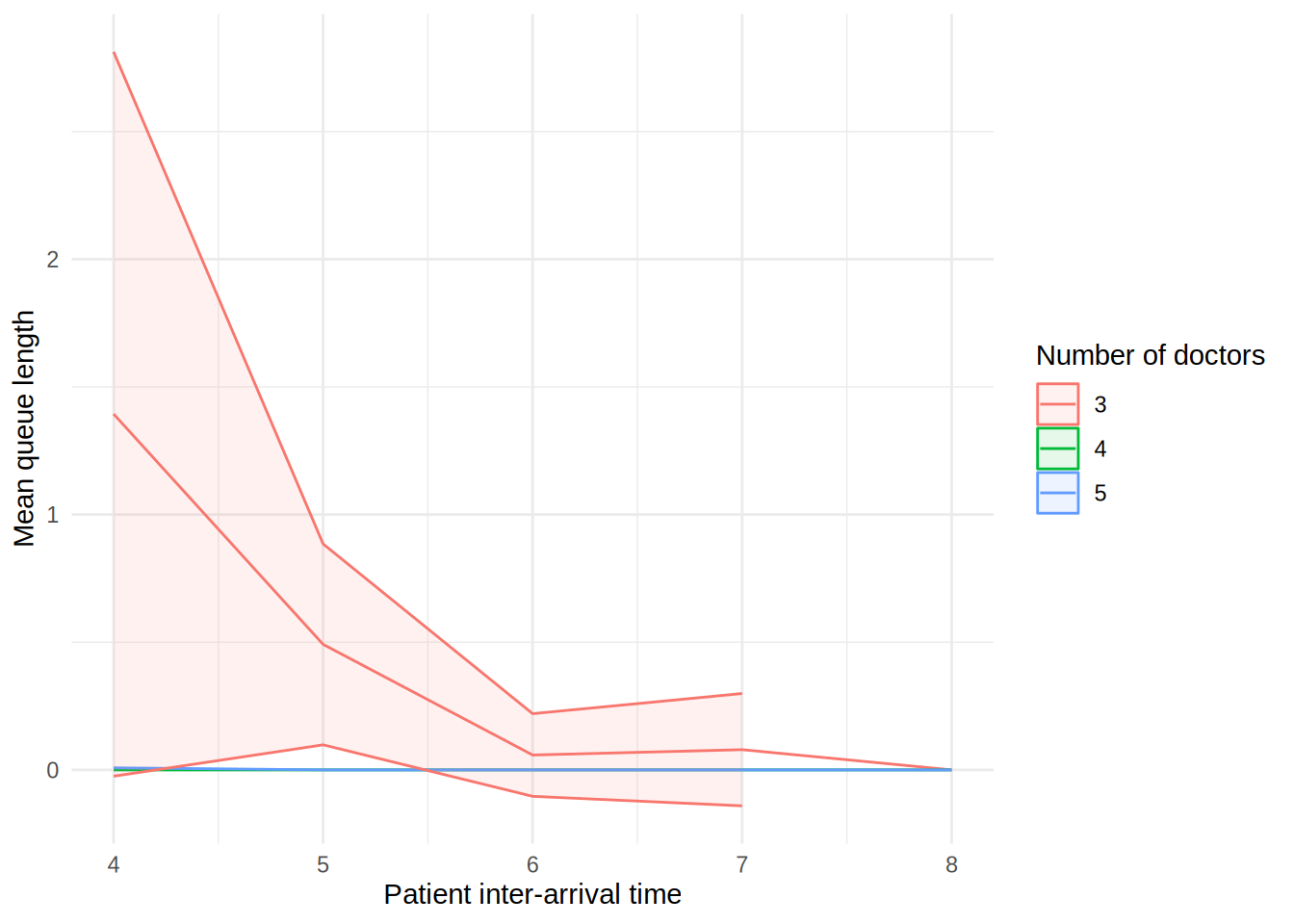
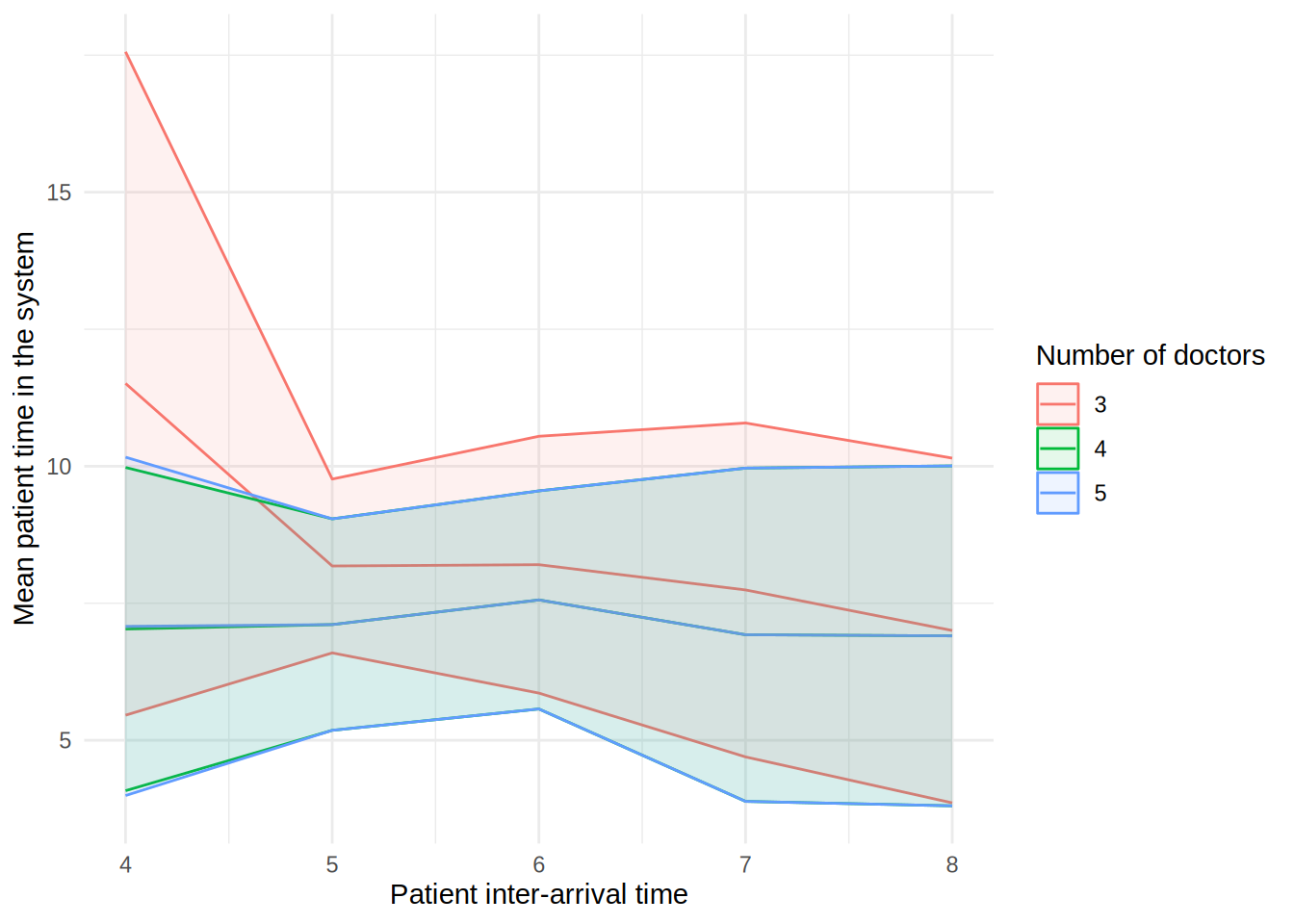
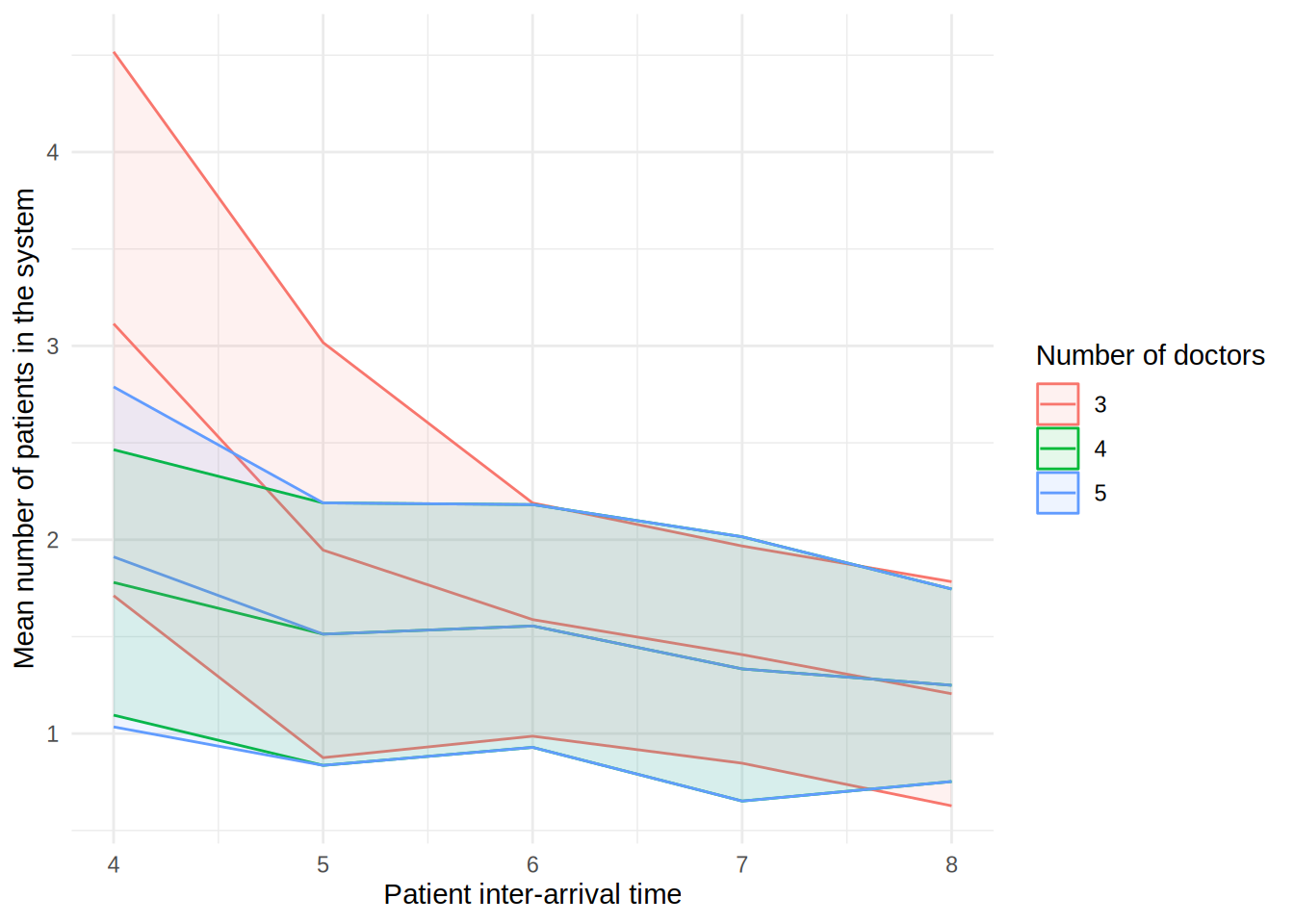
In the figure, each line shows the result (e.g., mean wait time for a doctor) as the patient inter-arrival time increases. The different coloured lines correspond to a different number of doctors. The shaded bands around each line represent 95% confidence intervals for the mean.
These plots can look a little funny simple due to the wide confidence intervals - with very short run times and few replications in our illustrative example, the confidence intervals are very wide!
- Use consistent names for variables on your axis labels.
- Include units in axis labels (e.g., minutes, days, patients per hour).
- Make sure to plot uncertainty (e.g., confidence intervals, percentile range).
- Keep colour and line style consistent across related figures.
Sensitivity analysis
sensitivity_plots = plot_metrics(
summary_tables=sensitivity_tables,
name_mappings=name_mappings,
path_prefix=os.path.join("tables_figures_resources", "python_sensitivity")
)Click the name of each metric below to view the relevant plot:
sensitivity_plots["mean_wait_time"]sensitivity_plots["mean_utilisation_tw"]sensitivity_plots["mean_queue_length"]sensitivity_plots["mean_time_in_system"]sensitivity_plots["mean_patients_in_system"]Sensitivity analysis
sensitivity_plots <- plot_metrics(
summary_tables = sensitivity_tables,
x_var = "interarrival_time",
name_mappings = name_mappings,
path_prefix = file.path("tables_figures_resources", "r_sensitivity")
)Click the name of each metric below to view the relevant plot:
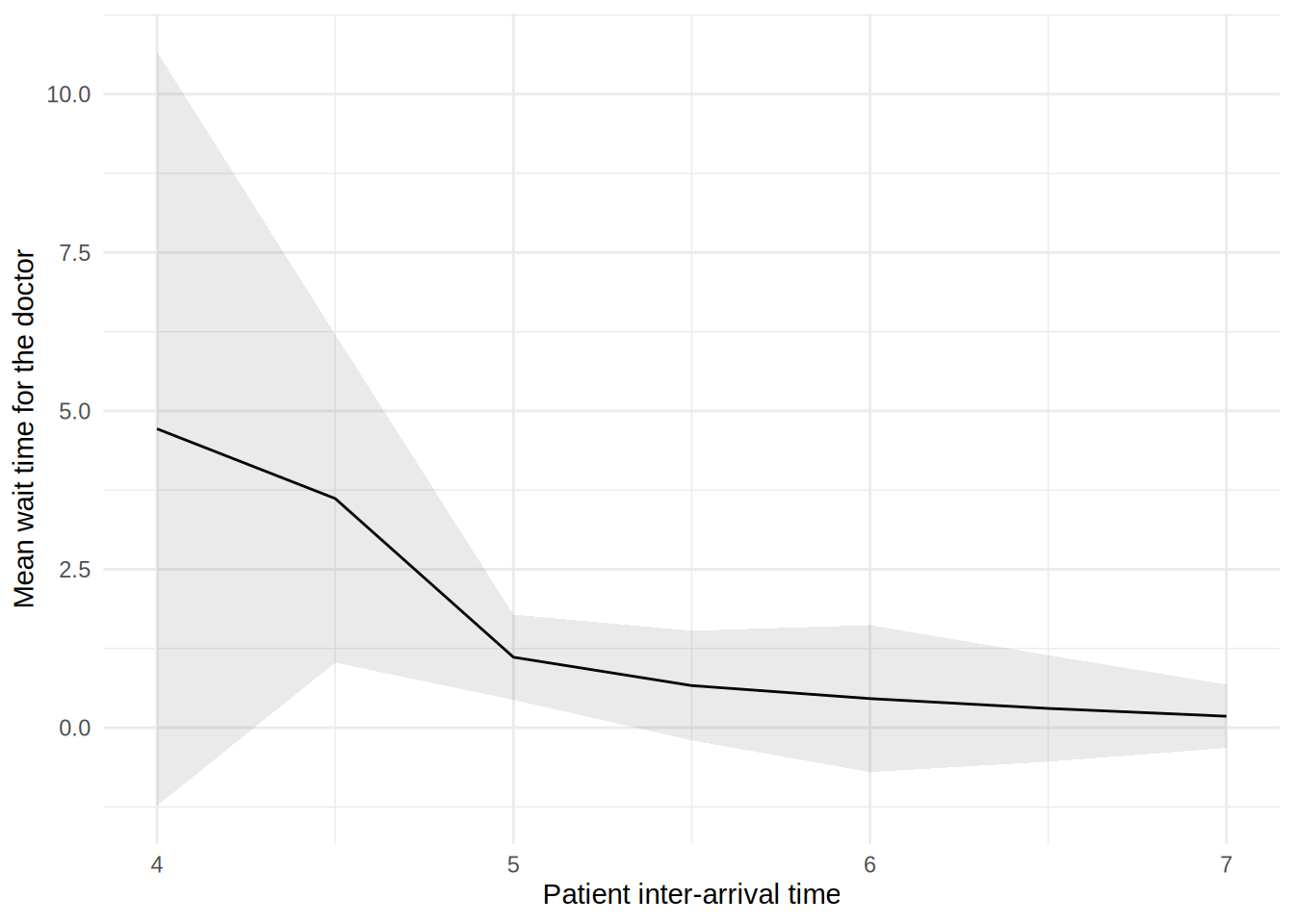
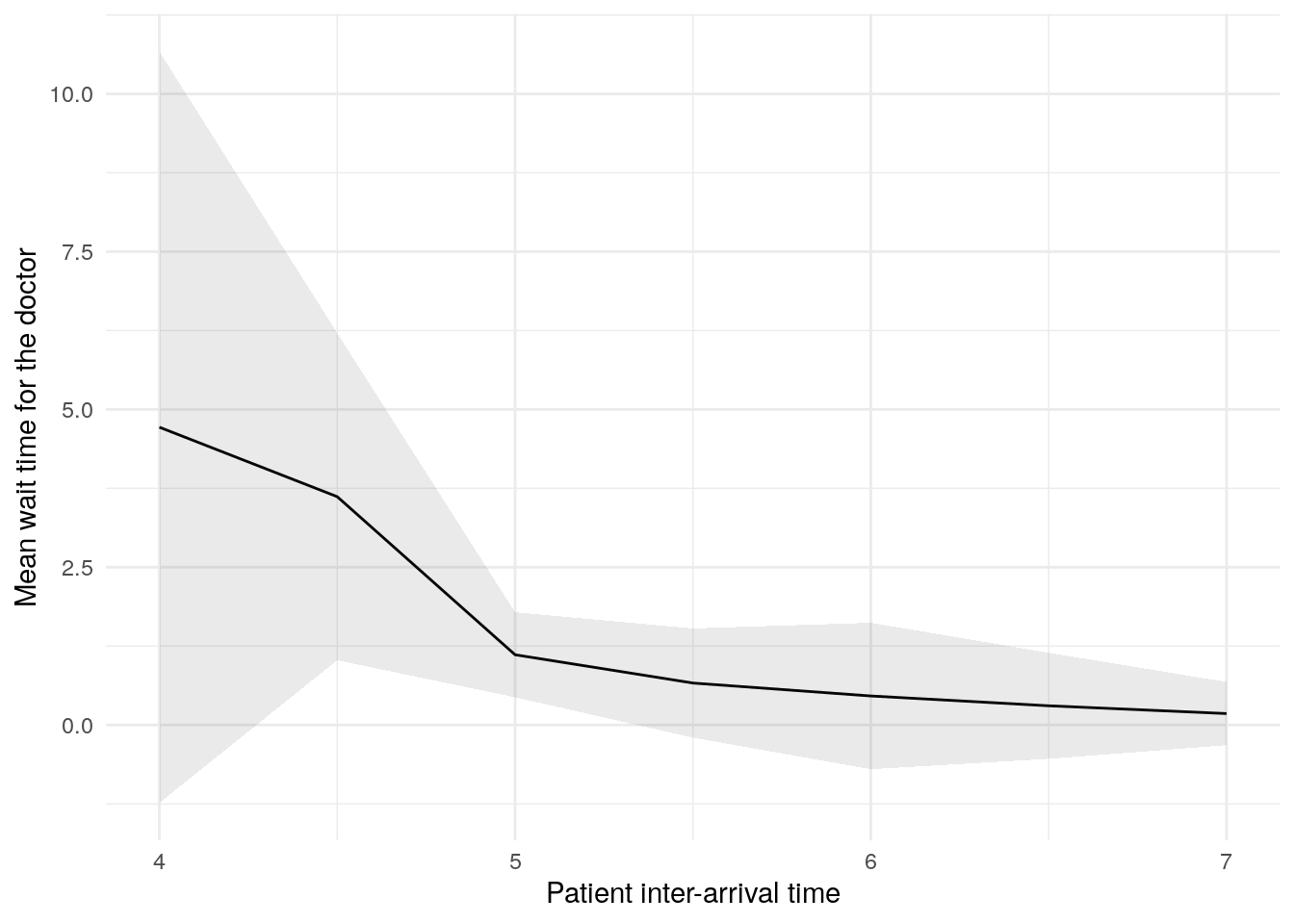
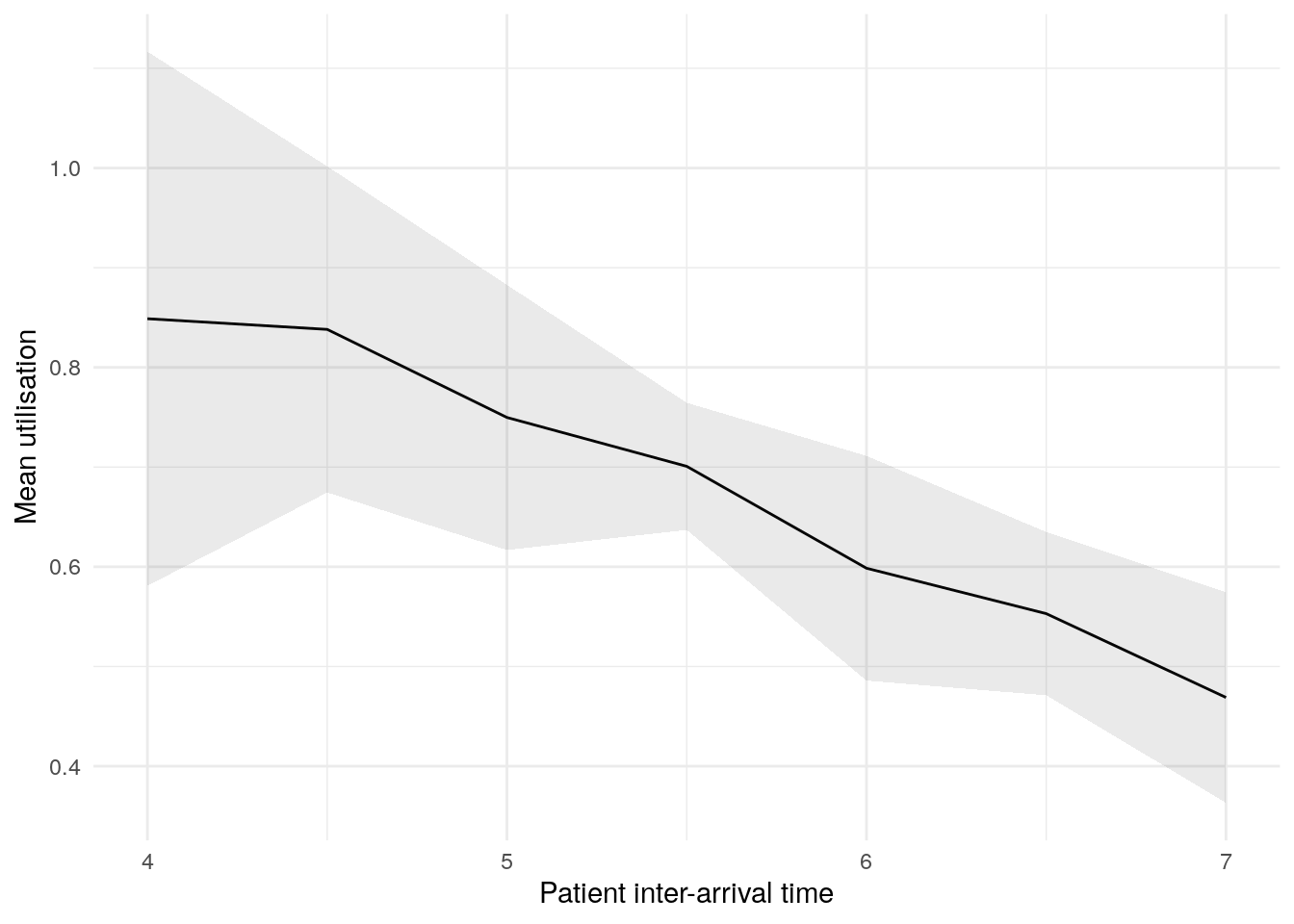
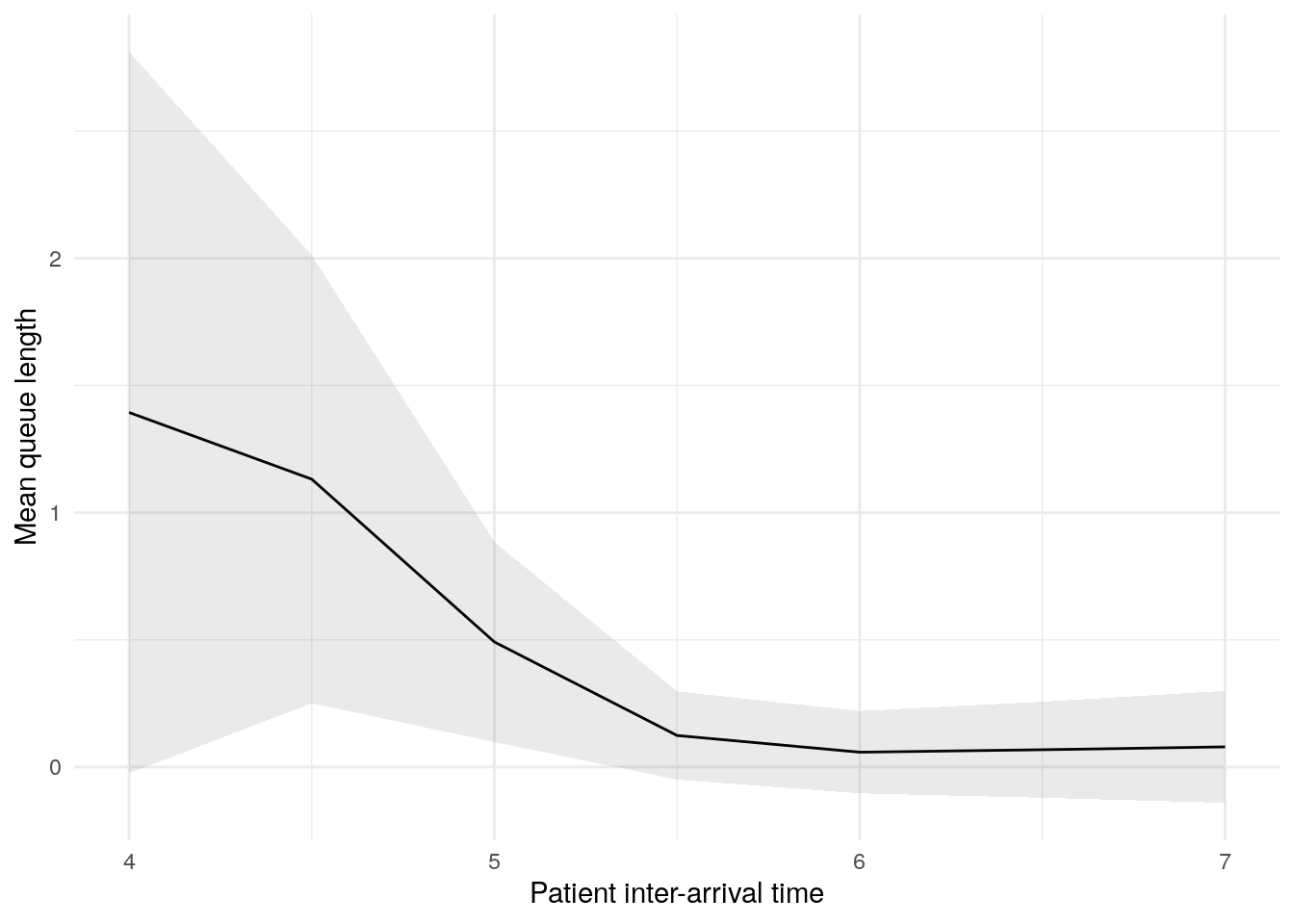
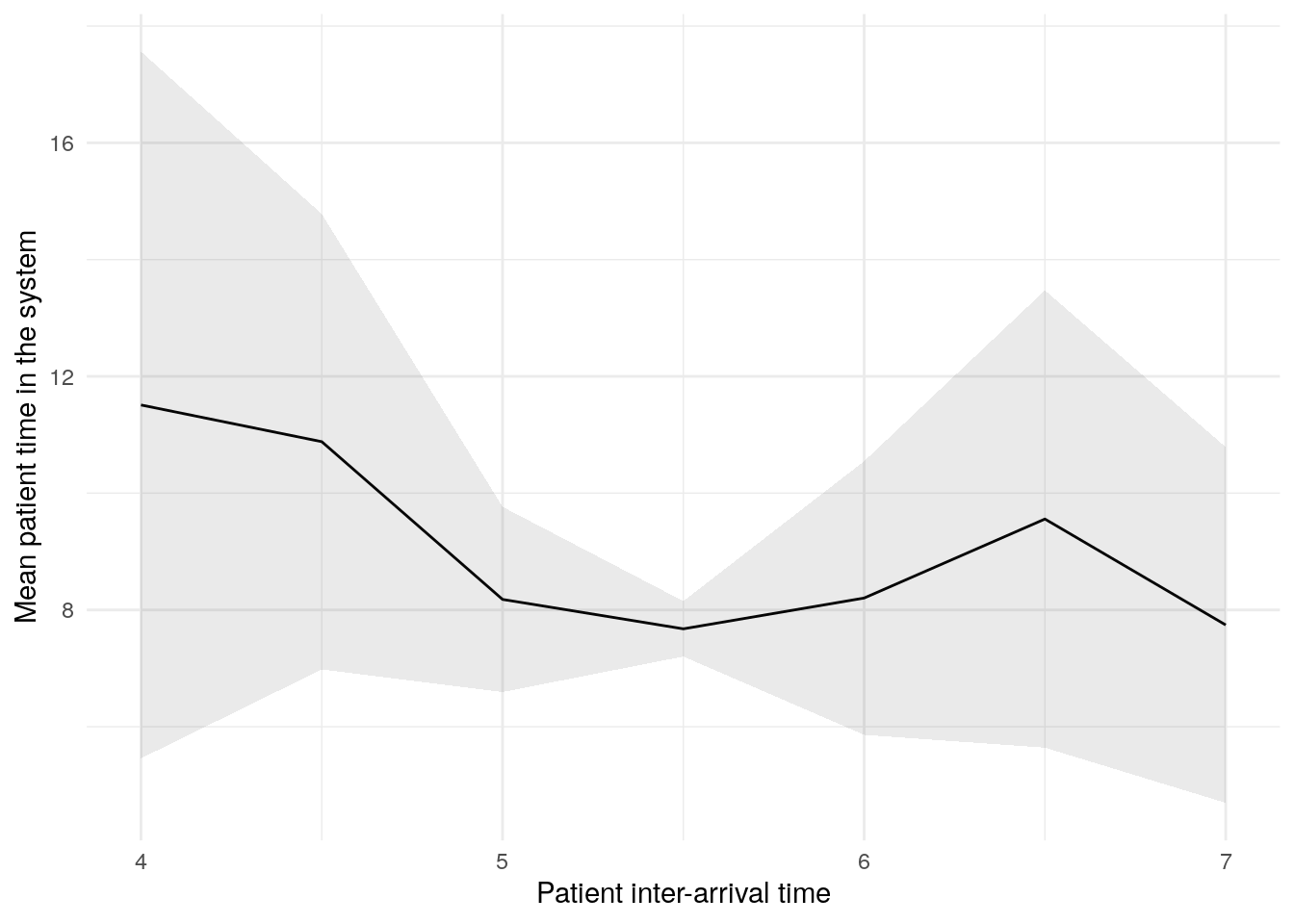
In this figure, the blue line shows the result (e.g., mean wait time) as we increase the patient inter-arrival time. The shaded band around the line is a 95% confidence interval, reflecting uncertainty due to random variation between simulation runs.
Explore the example models
Nurse visit simulation
Click to visit pydesrap_mms repository
| Key files | notebooks/analysis.ipynb |
Click to visit rdesrap_mms repository
| Key files | rmarkdown/analysis.md |
Stroke pathway simulation
Click to visit pydesrap_stroke repository
| Key files | notebooks/analysis.ipynb |
Click to visit rdesrap_stroke repository
| Key files | rmarkdown/analysis.md |
Test yourself
Quiz
Activity
Create at least one table and figure from your simulation results.
If you don’t have any to hand, feel free to download ours:
Make sure to write and save code that generates your outputs, and that you save the outputs to files (e.g., csv for tables, png for figures).
For example, create a Jupyter notebook for the analysis, and then push that to your GitHub repository.
For example, create an Rmarkdown file for the analysis, and then push that to your GitHub repository.